Status of Master Praktikum: open
Supervisors: Paolo Rota, Michael Reiter and Florian Kleber
Problem Statement
Acute Lymphoblastic Leukaemia (ALL) is the most frequent leukaemia entity in children and adolescents. Despite continued progress and refinement of therapeutic approaches, disease relapse due to insufficient extinction of leukaemic blasts still remains the number one cause of treatment failure.
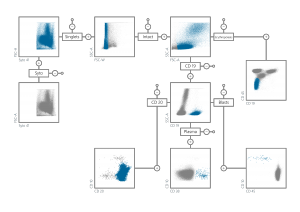
Flow cytometry (FCM) is one of the methodologies most useful in this respect, because it is widely available and applicable to most patients. A flow cytometer analysis single cells regarding physical (cell size and granularity) and biological properties (cell type). Based on the measurements, leukaemic cells are identified: the Minimal Residual Disease (MRD) value refers to the number of leukaemic cells in a blood sample. Currently this is done by a procedure called gating, where in 2D scatter plots different cell populations are labelled manually by polygons in a hierarchical way (see Figure). The data analysis rely largely on operator skills and experience. Thus, automated methods are needed for an objective MRD determination.
See also AutoFLOW project website.
Goal
Within the AutoFLOW project we set up a database consisting of FCM-MRD datasets with class labels for the most important cell populations from three dfferent time-points during chemotherapy from 199 patients with B-cell precursor ALL. This database should be used to test the R/BioConductor package/methodology flowDensity:
http://www.bioconductor.org/packages/release/bioc/html/flowDensity.html
and openCyto:
http://bioconductor.org/packages/release/bioc/html/openCyto.html
Thus, an interface to the current data structure must be defined, and the automated gating methods applied to the FCM data.
Workflow
- Literature research
- Get familiar with FCM data and R
- Development and evaluation of the system
- Written thesis (in English) and final presentation
Requirements
- R Project for Statistical Computing
References
http://www.bioconductor.org/packages/release/bioc/html/flowDensity.html
http://bioconductor.org/packages/release/bioc/html/openCyto.html
- K. Lo, R. Brinkman and R. Gottardo, “Automated gating of flow cytometry data via robust model based clustering”, Cytometry Part A, 73(4): 321-332.
- Bashashati and R. Brinkman, “A survey of flow cytometry data analysis methods.”, Advances in Bioinformatics, Article ID 584603, pp.1-19, 2009.
- N. Aghaeepour, G. Finak, H. Hoos, T. R. Mosmann, R. Brinkman, R. Gottardo, and R. H. Scheuermann, “Critical assessment of automated flow cytometry data analysis techniques.”, Nature Methods, vol. 10, no. 3, pp. 228–238, Mar. 2013.